Single-Cell Multi-Omics Experimental Design: How to Avoid Common Sample Preparation Mistakes?
Introduction
Single-cell multi-omics can reveal cell states, heterogeneity, and regulatory links at unprecedented resolution. But the results are only as good as the sample prep. If cells are stressed, contaminated, or poorly handled, the downstream data will be biased before sequencing even starts. This is why “how to do” sample preparation correctly matters more than any later analysis step. In this article, we focus on the most common mistakes in single-cell multi-omics experimental design and how to avoid them.
1. Why Sample Preparation Determines Single-Cell Data Quality
1.1 Contamination Control Starts Before Cell Capture
In single-cell workflows, contamination is not a minor technical issue. It can destroy library quality, distort cell profiles, and waste a full run. The core principle of aseptic technique is simple: build a barrier between the external microbial environment and the cells. That barrier must exist at every step, from the bench to the biosafety cabinet, from reagent handling to culture inspection.
Before any prep work begins, the workspace should be cleaned again, even if someone else already wiped it. Use 70% ethanol, then UV exposure for at least 30 minutes when appropriate. Keep only the materials needed for the current experiment inside the cabinet. Do not crowd the workspace. Do not let your hands or tools pass over open vessels. These are basic habits, but they are often the difference between usable and unusable data.
1.2 Cell Stress Is a Hidden Source of Bias
Single-cell multi-omics is especially sensitive to pre-analytical stress. Cells that are over-trypsinized, kept too long outside controlled conditions, or repeatedly exposed to temperature shifts may show altered expression before capture. That means the “biology” you measure may partly reflect handling artifacts.
A practical rule is to minimize time, force, and exposure. Work quickly, but do not rush. Every step should be planned before the sample enters the cabinet. If a sample drops, dries out, or spends too long waiting, the damage may be irreversible.
2. Common Mistakes in Sample Preparation
2.1 Poor Aseptic Practice
One of the most common errors is assuming the workspace is clean enough. It is not enough to trust the previous user’s cleaning. Re-disinfect the bench. Wear lab coat, mask, and sterile gloves. Long hair should be tied back. Gloves should overlap the lab coat cuffs.
If a reagent, plate, or tube leaves the biosafety cabinet, it should be re-cleaned before returning. Surfaces that were moved to a refrigerator, incubator, or sink must be wiped and disinfected again. The rule is simple: once a material leaves the sterile field, treat it as potentially contaminated.
Also avoid talking too much during open handling. Even minor airflow and droplets can increase contamination risk. In single-cell work, small errors scale into big losses.
2.2 Reagent and Medium Handling Errors
Reagents and media are frequent failure points. External packaging should be removed outside the cell room, and the outer surface should be disinfected with 70% ethanol before the container enters the work area. If the reagent is stored or moved outside the cabinet, clean it again before reuse.
For homemade solutions, label everything clearly. Include name, date, preparation details, and sterilization status. A well-labeled reagent is not administrative formality. It is experimental traceability. If a batch fails, precise labels help identify whether the issue came from the formulation, storage time, or handling.
Aliquot sensitive reagents when possible. Repeated freeze-thaw cycles damage enzymes and increase contamination risk. This is especially important when your workflow depends on multiple enzymatic steps.
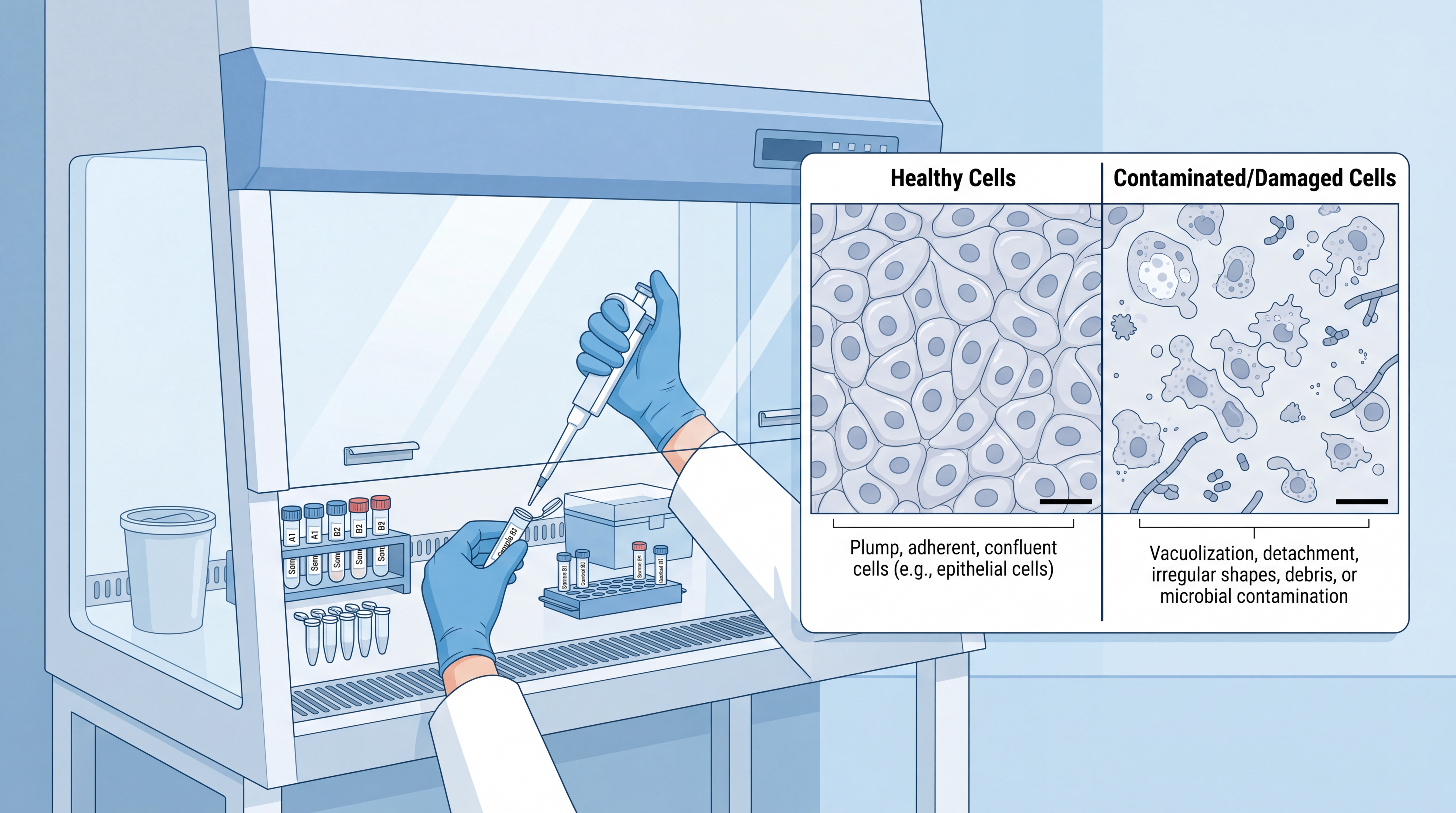
2.3 Incorrect Sample Quality Screening
Before any single-cell experiment, inspect the culture or tissue-derived suspension visually and under a microscope. Do not assume the sample is clean. If there is visible contamination, stop the run immediately and discard the affected material.
For cultured cells, a portion of each batch can be maintained separately in antibiotic-free complete medium as a contamination check. This helps reveal hidden problems that antibiotics may mask. In practice, this step is valuable because antibiotic use can delay detection, but it does not solve the underlying issue.
For sample preparation in multi-omics, quality screening should also include cell viability, aggregation, debris load, and the presence of dead cells. If the suspension is dirty or clumped, downstream capture efficiency will fall.
3. How to Build a Reliable Sample Preparation Workflow
3.1 Standardize the Work Area
A reliable workflow starts with a standardized cabinet setup. Only essential tools should be inside. Arrange them by function and habit, not randomly. Keep all actions within your line of sight. If you cannot fully see the tool or tip, you increase the chance of touching the tube rim, the bench, or another object.
If liquid spills on the bench, stop immediately and wipe with 70% ethanol. Clean the workspace again at the end of the session. This is not optional. It protects both the current sample and the next one.
3.2 Protect the Operator and the Sample
Personal protection is also sample protection. Wear gloves, mask, and lab coat correctly. If you are ill, such as with a cold, do not proceed casually. Either postpone the experiment or use stronger protection, such as a face shield, depending on institutional policy.
Minimize speaking near open samples. Keep gloves clean by wiping them with 70% ethanol at regular intervals. This matters because gloves contact multiple surfaces and can silently transfer contaminants. Good sample preparation is not only about technique. It is about disciplined behavior.
3.3 Plan the Sample Amount and Replicates Carefully
Single-cell multi-omics projects often fail because the design is underpowered or too complicated. For robust statistics, at least three biological replicates are needed. In animal studies, more are usually required because of individual variation, sex, age, and pathology-related bias.
A common practical principle is to keep the design simple and interpretable. Too many groups can create confusing comparisons and reduce the clarity of the biological question. The same logic applies to sample prep: do not overcomplicate the workflow if the sample quantity is limited or unstable.
If multiple batches are unavoidable, treat each batch as an independent data point during planning. Do not assume different batches are directly mergeable without considering variance and batch effects.
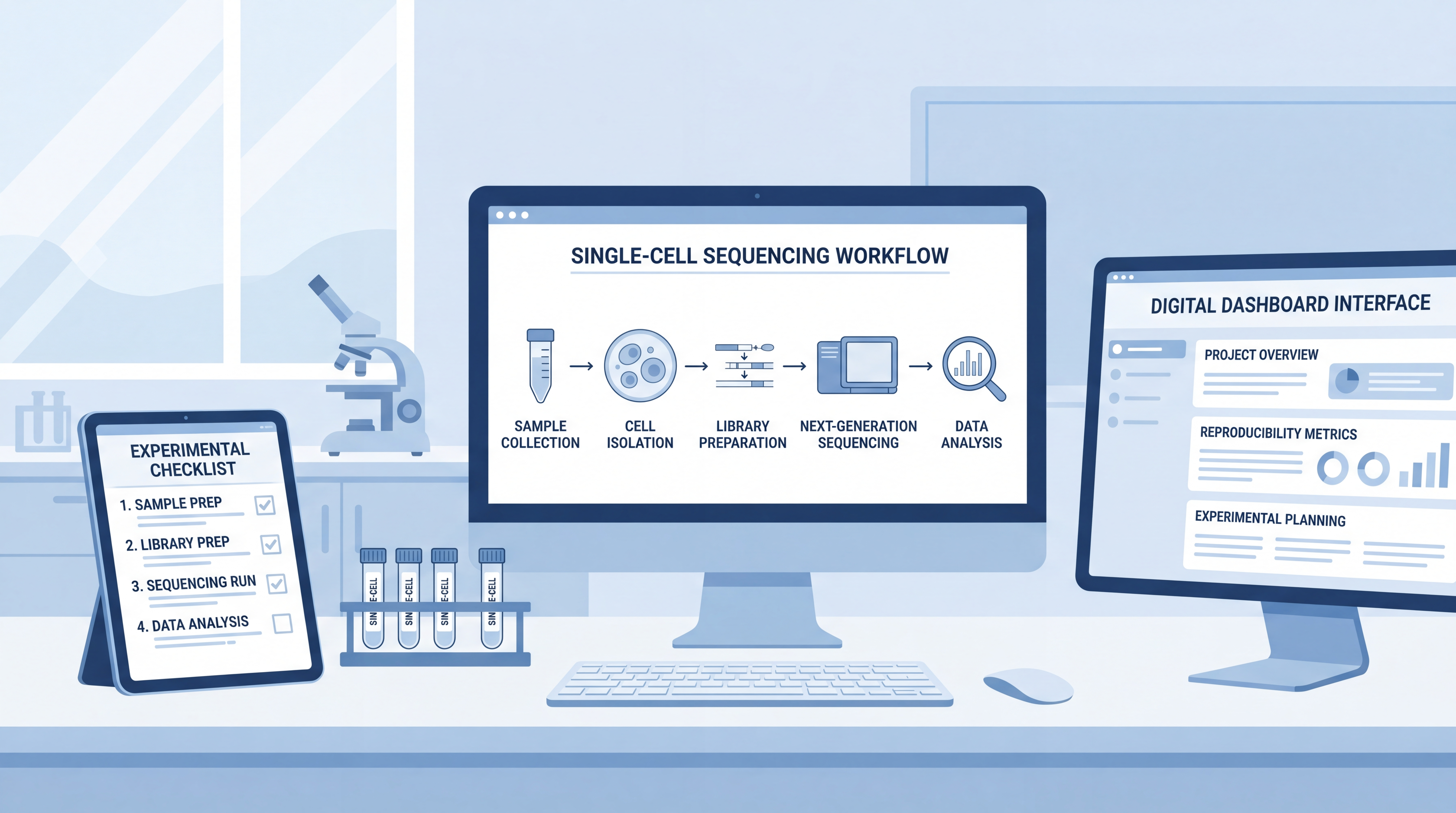
4. Practical Checklist Before Sequencing
4.1 Confirm Sample Integrity
Before loading the sample into a single-cell platform, confirm the following:
- No visible contamination.
- Acceptable cell viability.
- Low debris.
- Minimal clumping.
- Correct buffer composition.
- Proper temperature control.
If any of these fail, fix the problem before proceeding. Do not “push through” with a compromised sample. In multi-omics, bad input usually means bad output.
4.2 Keep the Workflow Consistent
Consistency improves reproducibility. Use the same preparation order, the same reagent sources when possible, and the same handling rules across samples. This reduces variation that has nothing to do with biology.
If a protocol includes mixing shared components, prepare a master mix first, then distribute it to each sample before adding variable components. This approach reduces pipetting inconsistency and helps maintain stable experimental conditions.
4.3 Use Digital Planning to Reduce Human Error
A major source of sample prep failure is not lack of skill. It is lack of planning. Tools such as scifocus.ai can help researchers structure workflows, organize experimental variables, and keep sample-handling plans clear. For complex single-cell multi-omics projects, that kind of organization saves time and reduces avoidable mistakes.
If your goal is to reduce failed runs and improve reproducibility, a planning layer like scifocus.ai can make your experimental design more disciplined and traceable. It supports better coordination from sample setup to downstream interpretation.
Conclusion
Single-cell multi-omics is powerful, but only if sample preparation is controlled. The main risks are contamination, cell stress, reagent handling errors, poor screening, and weak planning. The solution is straightforward: work aseptically, inspect samples carefully, label and store reagents properly, keep the workflow simple, and design with enough replication.
In practice, the quality of your data is decided long before sequencing begins. If you want fewer failures and cleaner experimental logic, adopt a structured planning process and use scifocus.ai to support your workflow design. That is the most practical way to reduce common sample preparation mistakes and move your project forward with confidence.
Did you like this article? Explore a few more related posts.
Start Your Research Journey With Scifocus Today
Create your free Scifocus account today and take your research to the next level. Experience the difference firsthand—your journey to academic excellence starts here.
